Cell Guidance Systems’ array genomic hybridisation (AGH) service provides high-resolution detection of copy number variations (CNVs) and loss of heterozygosity (LOH) in human cell lines, including iPSCs and ESCs. Using the Infinium Global Screening Array v3.0 with over 750,000 probes, AGH identifies acquired genomic abnormalities at a resolution of approximately 200 kb — far beyond what G-banding karyotyping alone can detect. We recommend combining AGH with karyotype analysis for a comprehensive genomic integrity assessment, as each method detects different classes of abnormality. Results are returned within 15–17 business days.






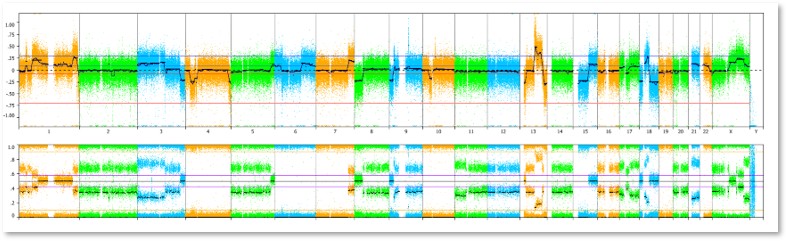
What types of abnormalities are reported?
Acquired copy number variations (CNVs) and loss of heterozygosity (LOH) present in at least ~15 – 20% of cells will be reported.
For the AGH analysis, an assessment of normal variation is made with reference to ~5,000 normal control samples and a database of ~10,000 clinical samples. However, benign constitutional (heritable) CNVs will not be reported unless requested.
Balanced rearrangements and low level (10 – 20%) of mosaicism will not be detected using AGH. In practice, this level of mosaicism is statistically similar to the level which can be detected by karyotype analysis of 20 cells.
We recommend array analysis for identifying marker chromosomes and additional material on a chromosome of unknown origin. Karyotype analysis alone can detect certain acquired abnormalities, e.g. gain in the small arm of a chromosome, but the precise nature of certain unbalanced chromosomal abnormalities is difficult to establish by only using G-banding analysis.
For more details, and a sample report see Services Description
Please note that we do not offer clinical services for patients.