Are free cytokines responsible for some of exosomes effects?

A recent review paper in Frontiers in Immunology highlights the possibility that free cytokine contaminants in isolated exosome samples may actually be causing biological effects that have been attributed to exosomes in immunomodulation studies.
Exosomes and cytokines share some features in common. Both are secreted by virtually all cells and enable cell-to-cell communication. Exosomes are more complex, larger structures that can incorporate proteins either into their walls or lumens.
Cytokines (a subset of growth factors) allow communication between cells of the immune system. The activity of the immune system is largely orchestrated by macrophages which secrete large numbers of exosomes and cytokines. Many cytokines are incorporated into exosomes for secretion. The proportion that is incorporated seems to depend on the type of cell and can vary depending on local conditions. For example, cytokine secretion profiles are dependent on the polarization state of macrophages.
A wide range of techniques has been developed to isolate/purify exosomes. These include ultracentrifugation, precipitation, affinity capture, size exclusion chromatography (SEC), and microfluidics. A perfect isolation technology (which captures all exosomes and excludes all other components) does not exist. However, there are distinct differences when it comes to purity. Ultimately, the method chosen for exosome isolation depends on historic precedent, sample volume, sample throughput requirements, and downstream applications. SEC is increasingly popular because it does most of these things well. For functional studies, there is evidence that SEC outperforms ultracentrifugation.
Given the difficulty in obtaining completely pure exosomes, and the similarities between exosome and cytokine function, it's perhaps not too surprising that cytokines may actually be responsible for some of the biological effects of the exosome samples. In order to assess the impact of the purification technique on cytokine contamination, specific assays need to be performed which assess the amount of free cytokine within an exosomes sample. Measuring the quantity of a cytokine present in a sample can be achieved using an ELISA assay specific for each cytokine.
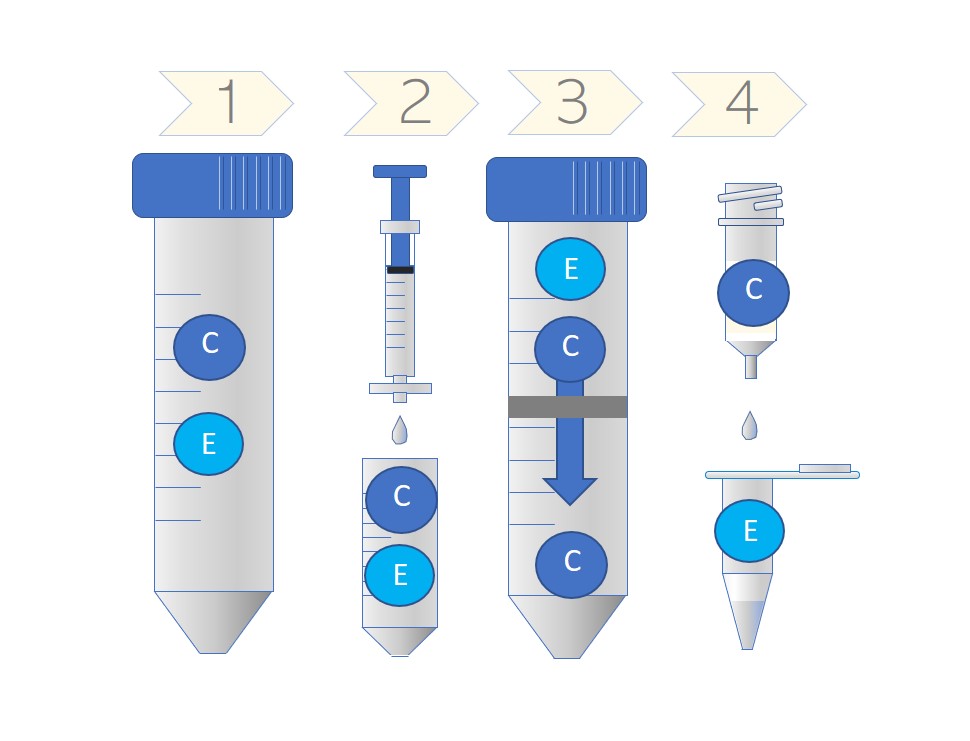
Schematic showing the co-purification or separation of cytokines (C) (and other smaller factors) and exosomes (E) at each step of the purification process (see main text below for details). In the final step, exosomes are almost completely separated from cytokines.
Shin La Shu and colleagues at the National Cancer Institute in Frederick and the Roswell Park Cancer Centre in Buffalo have found that the concentration of contaminating cytokines can be dramatically reduced dependent on the purification method used. Their process includes (1) removal of cells and cell debris by pelleting; (2) Filtering through a 0.2 micron filter to remove particles > 200 nm; (3) Concentration of the sample in a 100 kDa cut-off microconcentrator device (e.g. Vivaspin® or Amicon® and (4) purification using Exo-spin™ (an SEC column) to remove any remaning small particles and proteins including cytokines. Using this protocol, the cytokines and other proteins are primarily removed in the final two steps.
Addition of the microconcentrator step, in particular, generated higher purity reducing the levels of a range of cytokines by about an order of magnitude. In comparison with ultracentrifugation, the results were even more dramatic with an >800-fold reduction across a range of 13 cytokines measure by ELISA including IL-7, IL-8, IL-10 and IFN-γ. The authors didn’t compare the alternative concentration step 3 (using Exo-spin™ precipitant)
The protocol was also shown to provide higher yields compared to other techniques with a 25 to 58-fold higher yield than ultracentrifugation or ExoQuick™ Ultra respectively. This begs the question: what other contaminants does ultracentrifugation leave behind which are impacting the interpretation of downstream experiments? Moreover, when the MISEV requirements are updated, should this level of characterization be required for each sample?
IMAGE: Shutterstock